Home
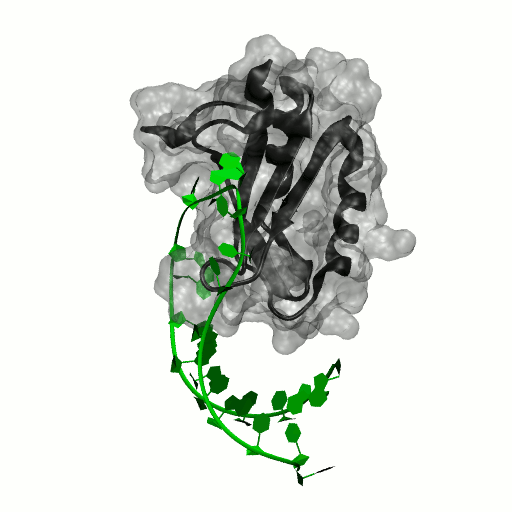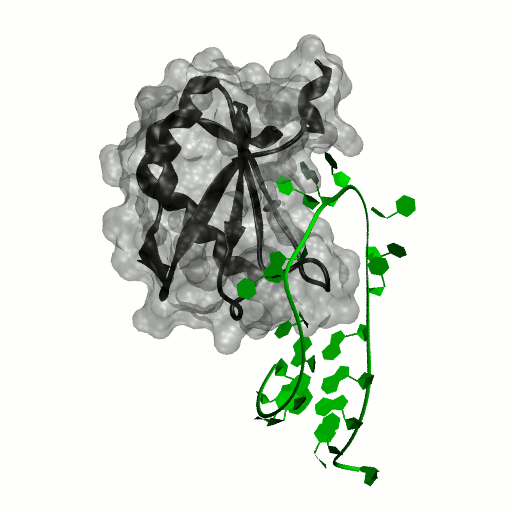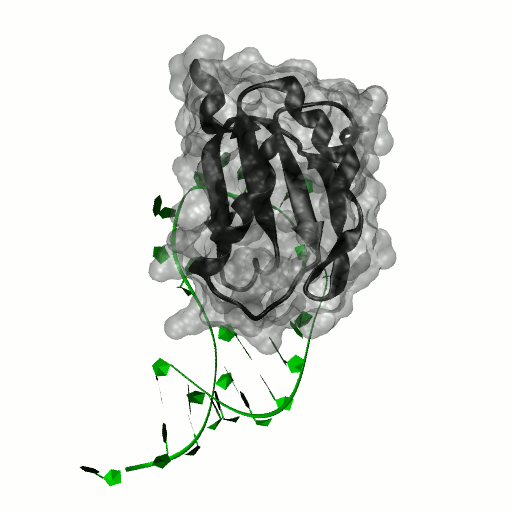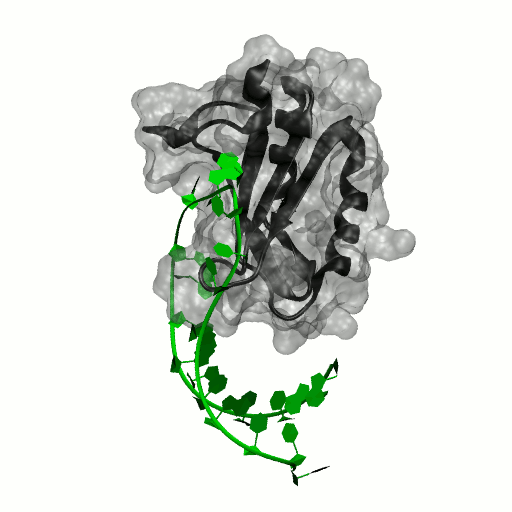
Structure by Oubridge et al 1994 (PDB: 1urn).
Click here to read the official Research Interests Book entry for Kathleen Hall.
RNA and RNA:Protein interactions provide biological regulation at all levels of cellular function. RNA molecules and their interactions with protein partners in order to understand their biology. We use biochemistry, molecular biology, and molecular biophysics to understand the properties of these molecules.
Which RNAs do we study?
We are interested in RNA structure, dynamics, and folding, and are currently investigating the properties of the Varkud Satellite Virus Ribozyme Stemloop V and the 60 nucleotide fragment of 23S rRNA that binds the L11 protein.
VSR SLV was first described by the Legault lab in 2005, using solution NMR methods. We are revisiting its structure and dynamics using NMR, fluorescence, and computational methods. This five nucleotide loop is a variant of a U-turn structure that is a common motif in RNAs, and as such it has some noncanonical features. This is a collaboration with the Cheatham lab (University of Utah).
The 60 nucleotide RNA from bacterial 23S rRNA that we study by NMR, steady-state and time-resolved fluorescence, stop-flow fluorescence, and computational methods has been the subject of several previous studies. Most notably, the Draper lab described the Mg2+ dependence of its tertiary folding and the thermodynamics of this association, and solved a crystal structure that shows it bound to a single K+ ion. The Schwalbe lab has studied its binding to both the L11 protein and to thiostrepton using NMR. Now, we are using specific 2-aminopurine labeling and specific 13C-NTP incorporation to probe its folding pathways, extracting folding kinetics, sites of Mg2+ association by NMR, and site-specific dynamics of its loops and junctions by NMR relaxation. Computational experiments probe K+ and Mg2+ ion association and exchange. This is also a collaboration with the Cheatham lab.
Which RNA:Protein interactions do we study?
Our work has focused for some time on human U1A protein and its recognition of two RNAs. In the U1 snRNP, U1A is bound to Stemloop II of the human U1 snRNA, where it uses seven out of ten nucleotides in the loop for specific recognition. In the nucleus, U1A also binds to its own pre-mRNA, in an interaction with the Polyadenylation Inhibition Element (PIE) RNA. This structure, found at the 3′ untranslated region of the U1A pre-mRNA, is bound by two U1A molecules to block adenylation. Only the N-terminal RNA Binding Domain (RBD) or RNA Recognition Motif (RRM) of the protein binds RNA; the function of the C-terminal RRM remains unknown.
The Drosophila SNF protein comes from a different branch of the U1A/U2B”/SNF evolutionary tree. The fly doesn’t have U1A protein, and it also lacks the related U2B” protein. Instead, it has SNF (sans fille). It binds both the U1 snRNA Stemloop II and the U2 snRNA Stemloop IV, and also binds the U2A’ protein. When Stemloop IV is bound to SNF, the affinity for U2A’ protein is enhanced, and when U2A’ is bound to SNF, the affinity for SLIV is increased. This example of heterotropic linkage is now the subject of thermodynamic, NMR, and computational investigations.